Project flow#
LaminDB allows tracking data lineage on the entire project level.
Here, we walk through exemplified app uploads, pipelines & notebooks following Schmidt et al., 2022.
A CRISPR screen reading out a phenotypic endpoint on T cells is paired with scRNA-seq to generate insights into IFN-γ production.
These insights get linked back to the original data through the steps taken in the project to provide context for interpretation & future decision making.

More specifically: Why should I care about data flow?
Data flow tracks data sources & transformations to trace biological insights, verify experimental outcomes, meet regulatory standards, increase the robustness of research and optimize the feedback loop of team-wide learning iterations.
While tracking data flow is easier when it’s governed by deterministic pipelines, it becomes hard when it’s governed by interactive human-driven analyses.
LaminDB interfaces workflow mangers for the former and embraces the latter.
Setup#
Init a test instance:
!lamin init --storage ./mydata
Show code cell output
💡 connected lamindb: testuser1/mydata
Import lamindb:
import lamindb as ln
from IPython.display import Image, display
💡 connected lamindb: testuser1/mydata
Steps#
In the following, we walk through exemplified steps covering different types of transforms (Transform).
Note
The full notebooks are in this repository.
App upload of phenotypic data  #
#
Register data through app upload from wetlab by testuser1:
# This function mimics the upload of artifacts via the UI
# In reality, you simply drag and drop files into the UI
def mock_upload_crispra_result_app():
ln.setup.login("testuser1")
transform = ln.Transform(name="Upload GWS CRISPRa result", type="upload")
ln.track(transform=transform)
output_path = ln.core.datasets.schmidt22_crispra_gws_IFNG(ln.settings.storage)
output_file = ln.Artifact(
output_path, description="Raw data of schmidt22 crispra GWS"
)
output_file.save()
mock_upload_crispra_result_app()
Show code cell output
💡 saved: Transform(uid='gp2akvy4K45UApul', name='Upload GWS CRISPRa result', type='upload', updated_at=2024-04-22 10:44:19 UTC, created_by_id=1)
💡 saved: Run(uid='3HuKvezDwkoSqk16Ummm', transform_id=1, created_by_id=1)
Hit identification in notebook  #
#
Access, transform & register data in drylab by testuser2:
# This function mimics the hit identification notebook
# In reality, you would run this in a notebook titled "GWS CRIPSRa analysis"
def mock_hit_identification_notebook():
# log in as another user
ln.setup.login("testuser2")
# create a new transform to mimic a new notebook
# (in reality you just run ln.track() in a notebook and you don't have to manage runs)
transform = ln.Transform(name="GWS CRIPSRa analysis", type="notebook")
transform.save()
run = ln.Run(transform=transform)
run.save()
# access the upload artifact
input_file = ln.Artifact.filter(key="schmidt22-crispra-gws-IFNG.csv").one()
# identify hits
input_df = input_file.load(is_run_input=run).set_index("id")
output_df = input_df[input_df["pos|fdr"] < 0.01].copy()
# register hits in output artifact
ln.Artifact.from_df(output_df, description="hits from schmidt22 crispra GWS", run=run).save()
mock_hit_identification_notebook()
Inspect data flow:
artifact = ln.Artifact.filter(description="hits from schmidt22 crispra GWS").one()
artifact.view_lineage()
Sequencer upload  #
#
Upload files from sequencer via script chromium_10x_upload.py:
!python project-flow-scripts/chromium_10x_upload.py
💡 connected lamindb: testuser1/mydata
💡 saved: Transform(uid='qCJPkOuZAi9q5zKv', name='chromium_10x_upload.py', key='chromium_10x_upload.py', version='1', type='script', updated_at=2024-04-22 10:44:23 UTC, created_by_id=1)
💡 saved: Run(uid='SaQAOMhCKX4JINEX9x61', transform_id=3, created_by_id=1)
✅ saved transform.source_code: Artifact(uid='kDjEQzGxm0nAYE9v7Qkv', suffix='.py', description='Source of transform qCJPkOuZAi9q5zKv', version='1', size=474, hash='o-QoKgEZGxbk5oBtcAKoWw', hash_type='md5', visibility=0, key_is_virtual=True, updated_at=2024-04-22 10:44:23 UTC, storage_id=1, created_by_id=1)
✅ saved run.environment: Artifact(uid='AzhIU0kSdZK8VUG22FYN', suffix='.txt', description='requirements.txt', size=3400, hash='lrHLv26VOiHp8JkBy2B7zw', hash_type='md5', visibility=0, key_is_virtual=True, updated_at=2024-04-22 10:44:23 UTC, storage_id=1, created_by_id=1)
scRNA-seq bioinformatics pipeline  #
#
Process uploaded files using a script or workflow manager: Pipelines and obtain 3 output files in a directory filtered_feature_bc_matrix/:
!python project-flow-scripts/cellranger.py
💡 connected lamindb: testuser1/mydata
💡 saved: Transform(uid='9bNIXJwR225zDPfo', name='Cell Ranger', version='7.2.0', type='pipeline', reference='https://www.10xgenomics.com/support/software/cell-ranger/7.2', updated_at=2024-04-22 10:44:25 UTC, created_by_id=2)
💡 saved: Run(uid='wHEfJWmQON31skSN0Jx9', transform_id=4, created_by_id=2)
❗ this creates one artifact per file in the directory - you might simply call ln.Artifact(dir) to get one artifact for the entire directory
!python project-flow-scripts/postprocess_cellranger.py
💡 connected lamindb: testuser1/mydata
💡 saved: Transform(uid='YqmbO6oMXjRj65cN', name='postprocess_cellranger.py', key='postprocess_cellranger.py', version='2', type='script', updated_at=2024-04-22 10:44:27 UTC, created_by_id=2)
💡 saved: Run(uid='0A2CiHinjtaQGuvqykxu', transform_id=5, created_by_id=2)
✅ saved transform.source_code: Artifact(uid='7akxxWYex7HxOVrE6Hjn', suffix='.py', description='Source of transform YqmbO6oMXjRj65cN', version='2', size=495, hash='iLSbWXZ-j7pkIgzO0i6c0w', hash_type='md5', visibility=0, key_is_virtual=True, updated_at=2024-04-22 10:44:28 UTC, storage_id=1, created_by_id=2)
❗ returning existing artifact with same hash: Artifact(uid='AzhIU0kSdZK8VUG22FYN', suffix='.txt', description='requirements.txt', size=3400, hash='lrHLv26VOiHp8JkBy2B7zw', hash_type='md5', visibility=0, key_is_virtual=True, updated_at=2024-04-22 10:44:23 UTC, storage_id=1, created_by_id=1)
✅ saved run.environment: Artifact(uid='AzhIU0kSdZK8VUG22FYN', suffix='.txt', description='requirements.txt', size=3400, hash='lrHLv26VOiHp8JkBy2B7zw', hash_type='md5', visibility=0, key_is_virtual=True, updated_at=2024-04-22 10:44:23 UTC, storage_id=1, created_by_id=1)
Inspect data flow:
output_file = ln.Artifact.filter(description="perturbseq counts").one()
output_file.view_lineage()
Integrate scRNA-seq & phenotypic data  #
#
Integrate data in a notebook:
# This function mimics the integrated analysis notebook
# In reality, you would run this in a notebook titled "Perform single cell analysis, integrate with CRISPRa screen"
def mock_integrated_analysis_notebook():
import scanpy as sc
# Create a new transform to mimic a new notebook
# In reality you just run ln.track() in a notebook
transform = ln.Transform(
name="Perform single cell analysis, integrate with CRISPRa screen",
type="notebook",
)
transform.save()
run = ln.Run(transform=transform)
run.save()
# access the output files of bfx pipeline and previous analysis
file_ps = ln.Artifact.filter(description__icontains="perturbseq").one()
adata = file_ps.load(is_run_input=run)
file_hits = ln.Artifact.filter(description="hits from schmidt22 crispra GWS").one()
screen_hits = file_hits.load(is_run_input=run)
# perform analysis and register output plot files
sc.tl.score_genes(adata, adata.var_names.intersection(screen_hits.index).tolist())
filesuffix = "_fig1_score-wgs-hits.png"
sc.pl.umap(adata, color="score", show=False, save=filesuffix)
filepath = f"figures/umap{filesuffix}"
artifact = ln.Artifact(filepath, key=filepath, run=run)
artifact.save()
filesuffix = "fig2_score-wgs-hits-per-cluster.png"
sc.pl.matrixplot(
adata, groupby="cluster_name", var_names=["score"], show=False, save=filesuffix
)
filepath = f"figures/matrixplot_{filesuffix}"
artifact = ln.Artifact(filepath, key=filepath, run=run)
artifact.save()
mock_integrated_analysis_notebook()
Show code cell output
WARNING: saving figure to file figures/umap_fig1_score-wgs-hits.png
WARNING: saving figure to file figures/matrixplot_fig2_score-wgs-hits-per-cluster.png
Review results#
Let’s load one of the plots:
# track the current notebook as transform
ln.settings.transform.stem_uid = "1LCd8kco9lZU"
ln.settings.transform.version = "0"
ln.track()
💡 notebook imports: ipython==8.23.0 lamindb==0.70.3 scanpy==1.10.1
💡 saved: Transform(uid='1LCd8kco9lZU6K79', name='Project flow', key='project-flow', version='0', type='notebook', updated_at=2024-04-22 10:44:30 UTC, created_by_id=1)
💡 saved: Run(uid='82uYYFhqEFomf2JXDttQ', transform_id=7, created_by_id=1)
artifact = ln.Artifact.filter(key__contains="figures/matrixplot").one()
artifact.cache()
Show code cell output
PosixUPath('/home/runner/work/lamin-usecases/lamin-usecases/docs/mydata/.lamindb/oMKVSi1DRuemsvVBtQA1.png')
display(Image(filename=artifact.path))
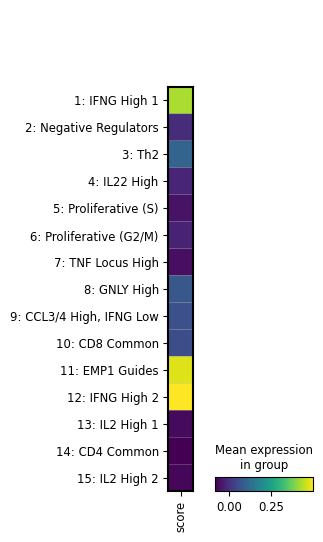
We see that the image artifact is tracked as an input of the current notebook. The input is highlighted, the notebook follows at the bottom:
artifact.view_lineage()
Alternatively, we can also look at the sequence of transforms:
transform = ln.Transform.search("Project flow", return_queryset=True).first()
transform.parents.df()
| uid | name | key | version | description | type | latest_report_id | source_code_id | reference | reference_type | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||
| 6 | UwZc0pDVwSJruSbZ | Perform single cell analysis, integrate with C... | None | None | None | notebook | None | None | None | None | 2024-04-22 10:44:30.132701+00:00 | 2024-04-22 10:44:30.132732+00:00 | 1 |
transform.view_parents()
Understand runs#
We tracked pipeline and notebook runs through run_context, which stores a Transform and a Run record as a global context.
Artifact objects are the inputs and outputs of runs.
What if I don’t want a global context?
Sometimes, we don’t want to create a global run context but manually pass a run when creating an artifact:
run = ln.Run(transform=transform)
ln.Artifact(filepath, run=run)
When does an artifact appear as a run input?
When accessing an artifact via stage(), load() or backed(), two things happen:
The current run gets added to
artifact.input_ofThe transform of that artifact gets added as a parent of the current transform
You can then switch off auto-tracking of run inputs if you set ln.settings.track_run_inputs = False: Can I disable tracking run inputs?
You can also track run inputs on a case by case basis via is_run_input=True, e.g., here:
artifact.load(is_run_input=True)
Query by provenance#
We can query or search for the notebook that created the artifact:
transform = ln.Transform.search("GWS CRIPSRa analysis", return_queryset=True).first()
And then find all the artifacts created by that notebook:
ln.Artifact.filter(transform=transform).df()
| uid | storage_id | key | suffix | accessor | description | version | size | hash | hash_type | n_objects | n_observations | transform_id | run_id | visibility | key_is_virtual | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||
| 2 | ONjKRMNVKarELEit8oE1 | 1 | None | .parquet | DataFrame | hits from schmidt22 crispra GWS | None | 18368 | PihzyuN-FWc-ld6ioxAuPg | md5 | None | None | 2 | 2 | 1 | True | 2024-04-22 10:44:21.151206+00:00 | 2024-04-22 10:44:21.151234+00:00 | 1 |
Which transform ingested a given artifact?
artifact = ln.Artifact.filter().first()
artifact.transform
Transform(uid='gp2akvy4K45UApul', name='Upload GWS CRISPRa result', type='upload', updated_at=2024-04-22 10:44:19 UTC, created_by_id=1)
And which user?
artifact.created_by
User(uid='DzTjkKse', handle='testuser1', name='Test User1', updated_at=2024-04-22 10:44:22 UTC)
Which transforms were created by a given user?
users = ln.User.lookup()
ln.Transform.filter(created_by=users.testuser1).df()
| uid | name | key | version | description | type | latest_report_id | source_code_id | reference | reference_type | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||
| 1 | gp2akvy4K45UApul | Upload GWS CRISPRa result | None | None | None | upload | None | NaN | None | None | 2024-04-22 10:44:19.706497+00:00 | 2024-04-22 10:44:19.706518+00:00 | 1 |
| 2 | fDSxN3HIA4HZlPlm | GWS CRIPSRa analysis | None | None | None | notebook | None | NaN | None | None | 2024-04-22 10:44:21.100198+00:00 | 2024-04-22 10:44:21.100224+00:00 | 1 |
| 3 | qCJPkOuZAi9q5zKv | chromium_10x_upload.py | chromium_10x_upload.py | 1 | None | script | None | 3.0 | None | None | 2024-04-22 10:44:23.064406+00:00 | 2024-04-22 10:44:23.534421+00:00 | 1 |
| 6 | UwZc0pDVwSJruSbZ | Perform single cell analysis, integrate with C... | None | None | None | notebook | None | NaN | None | None | 2024-04-22 10:44:30.132701+00:00 | 2024-04-22 10:44:30.132732+00:00 | 1 |
| 7 | 1LCd8kco9lZU6K79 | Project flow | project-flow | 0 | None | notebook | None | NaN | None | None | 2024-04-22 10:44:30.969187+00:00 | 2024-04-22 10:44:30.969214+00:00 | 1 |
Which notebooks were created by a given user?
ln.Transform.filter(created_by=users.testuser1, type="notebook").df()
| uid | name | key | version | description | type | latest_report_id | source_code_id | reference | reference_type | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||
| 2 | fDSxN3HIA4HZlPlm | GWS CRIPSRa analysis | None | None | None | notebook | None | None | None | None | 2024-04-22 10:44:21.100198+00:00 | 2024-04-22 10:44:21.100224+00:00 | 1 |
| 6 | UwZc0pDVwSJruSbZ | Perform single cell analysis, integrate with C... | None | None | None | notebook | None | None | None | None | 2024-04-22 10:44:30.132701+00:00 | 2024-04-22 10:44:30.132732+00:00 | 1 |
| 7 | 1LCd8kco9lZU6K79 | Project flow | project-flow | 0 | None | notebook | None | None | None | None | 2024-04-22 10:44:30.969187+00:00 | 2024-04-22 10:44:30.969214+00:00 | 1 |
We can also view all recent additions to the entire database:
ln.view()
Show code cell output
Artifact
| uid | storage_id | key | suffix | accessor | description | version | size | hash | hash_type | n_objects | n_observations | transform_id | run_id | visibility | key_is_virtual | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||
| 13 | oMKVSi1DRuemsvVBtQA1 | 1 | figures/matrixplot_fig2_score-wgs-hits-per-clu... | .png | None | None | None | 28814 | 8zXF_cVwaZnfhmrLbt_0kA | md5 | None | None | 6 | 6 | 1 | True | 2024-04-22 10:44:30.717957+00:00 | 2024-04-22 10:44:30.717985+00:00 | 1 |
| 12 | 1X7ygRzw9r9PYTZJSlWO | 1 | figures/umap_fig1_score-wgs-hits.png | .png | None | None | None | 118999 | DCFDLUMF-UohaBvkThn0mA | md5 | None | None | 6 | 6 | 1 | True | 2024-04-22 10:44:30.394167+00:00 | 2024-04-22 10:44:30.394193+00:00 | 1 |
| 11 | yjHnz6loDAgXJGxICSx0 | 1 | schmidt22_perturbseq.h5ad | .h5ad | AnnData | perturbseq counts | None | 20659936 | la7EvqEUMDlug9-rpw-udA | md5 | None | None | 5 | 5 | 1 | False | 2024-04-22 10:44:28.912912+00:00 | 2024-04-22 10:44:28.912944+00:00 | 2 |
| 9 | Ivw8X9xfLGngps3dF3Bu | 1 | perturbseq/filtered_feature_bc_matrix/matrix.m... | .mtx.gz | None | None | None | 6 | agsY6Sewjf8A4u971GjAgA | md5 | None | None | 4 | 4 | 1 | False | 2024-04-22 10:44:26.150419+00:00 | 2024-04-22 10:44:26.150437+00:00 | 2 |
| 8 | AtzCqKd7PCNSsZ2sKwtq | 1 | perturbseq/filtered_feature_bc_matrix/features... | .tsv.gz | None | None | None | 6 | 4tCaUwVhKL7NZ4m3okJUxw | md5 | None | None | 4 | 4 | 1 | False | 2024-04-22 10:44:26.149841+00:00 | 2024-04-22 10:44:26.149861+00:00 | 2 |
| 7 | gsBLhW9whA6D3dTKejBk | 1 | perturbseq/filtered_feature_bc_matrix/barcodes... | .tsv.gz | None | None | None | 6 | UrZgFbo_u8hv76AyDqDjQg | md5 | None | None | 4 | 4 | 1 | False | 2024-04-22 10:44:26.149021+00:00 | 2024-04-22 10:44:26.149044+00:00 | 2 |
| 6 | Hz3TM9GTNpiZnJedAzCm | 1 | fastq/perturbseq_R2_001.fastq.gz | .fastq.gz | None | None | None | 6 | QQoZzBXmdaLTtV0iPavwcQ | md5 | None | None | 3 | 3 | 1 | False | 2024-04-22 10:44:23.543689+00:00 | 2024-04-22 10:44:23.543708+00:00 | 1 |
Run
| uid | transform_id | started_at | finished_at | created_by_id | json | report_id | environment_id | is_consecutive | reference | reference_type | created_at | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | ||||||||||||
| 1 | 3HuKvezDwkoSqk16Ummm | 1 | 2024-04-22 10:44:19.710565+00:00 | NaT | 1 | None | None | NaN | True | None | None | 2024-04-22 10:44:19.710676+00:00 |
| 2 | ETwv5Fg5bCsY2ziKWnVG | 2 | 2024-04-22 10:44:21.101992+00:00 | NaT | 1 | None | None | NaN | None | None | None | 2024-04-22 10:44:21.102093+00:00 |
| 3 | SaQAOMhCKX4JINEX9x61 | 3 | 2024-04-22 10:44:23.066821+00:00 | 2024-04-22 10:44:23.545684+00:00 | 1 | None | None | 4.0 | None | None | None | 2024-04-22 10:44:23.066933+00:00 |
| 4 | wHEfJWmQON31skSN0Jx9 | 4 | 2024-04-22 10:44:25.678299+00:00 | NaT | 2 | None | None | NaN | None | None | None | 2024-04-22 10:44:25.678393+00:00 |
| 5 | 0A2CiHinjtaQGuvqykxu | 5 | 2024-04-22 10:44:27.787904+00:00 | NaT | 2 | None | None | 4.0 | None | None | None | 2024-04-22 10:44:27.788001+00:00 |
| 6 | vDSFxC4FmPbZDk1prQXS | 6 | 2024-04-22 10:44:30.135408+00:00 | NaT | 1 | None | None | NaN | None | None | None | 2024-04-22 10:44:30.135513+00:00 |
| 7 | 82uYYFhqEFomf2JXDttQ | 7 | 2024-04-22 10:44:30.974576+00:00 | NaT | 1 | None | None | NaN | True | None | None | 2024-04-22 10:44:30.974676+00:00 |
Storage
| uid | root | description | type | region | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|
| id | ||||||||
| 1 | ymLjnSrk | /home/runner/work/lamin-usecases/lamin-usecase... | None | local | None | 2024-04-22 10:44:17.864230+00:00 | 2024-04-22 10:44:17.864250+00:00 | 1 |
Transform
| uid | name | key | version | description | type | latest_report_id | source_code_id | reference | reference_type | created_at | updated_at | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||
| 7 | 1LCd8kco9lZU6K79 | Project flow | project-flow | 0 | None | notebook | None | NaN | None | None | 2024-04-22 10:44:30.969187+00:00 | 2024-04-22 10:44:30.969214+00:00 | 1 |
| 6 | UwZc0pDVwSJruSbZ | Perform single cell analysis, integrate with C... | None | None | None | notebook | None | NaN | None | None | 2024-04-22 10:44:30.132701+00:00 | 2024-04-22 10:44:30.132732+00:00 | 1 |
| 5 | YqmbO6oMXjRj65cN | postprocess_cellranger.py | postprocess_cellranger.py | 2 | None | script | None | 10.0 | None | None | 2024-04-22 10:44:27.785576+00:00 | 2024-04-22 10:44:28.249952+00:00 | 2 |
| 4 | 9bNIXJwR225zDPfo | Cell Ranger | None | 7.2.0 | None | pipeline | None | NaN | https://www.10xgenomics.com/support/software/c... | None | 2024-04-22 10:44:25.675861+00:00 | 2024-04-22 10:44:25.675885+00:00 | 2 |
| 3 | qCJPkOuZAi9q5zKv | chromium_10x_upload.py | chromium_10x_upload.py | 1 | None | script | None | 3.0 | None | None | 2024-04-22 10:44:23.064406+00:00 | 2024-04-22 10:44:23.534421+00:00 | 1 |
| 2 | fDSxN3HIA4HZlPlm | GWS CRIPSRa analysis | None | None | None | notebook | None | NaN | None | None | 2024-04-22 10:44:21.100198+00:00 | 2024-04-22 10:44:21.100224+00:00 | 1 |
| 1 | gp2akvy4K45UApul | Upload GWS CRISPRa result | None | None | None | upload | None | NaN | None | None | 2024-04-22 10:44:19.706497+00:00 | 2024-04-22 10:44:19.706518+00:00 | 1 |
User
| uid | handle | name | created_at | updated_at | |
|---|---|---|---|---|---|
| id | |||||
| 2 | bKeW4T6E | testuser2 | Test User2 | 2024-04-22 10:44:21.092451+00:00 | 2024-04-22 10:44:25.651998+00:00 |
| 1 | DzTjkKse | testuser1 | Test User1 | 2024-04-22 10:44:17.861305+00:00 | 2024-04-22 10:44:22.933146+00:00 |
Show code cell content
!lamin login testuser1
!lamin delete --force mydata
!rm -r ./mydata
✅ logged in with email testuser1@lamin.ai (uid: DzTjkKse)
💡 deleting instance testuser1/mydata
❗ manually delete your stored data: /home/runner/work/lamin-usecases/lamin-usecases/docs/mydata
